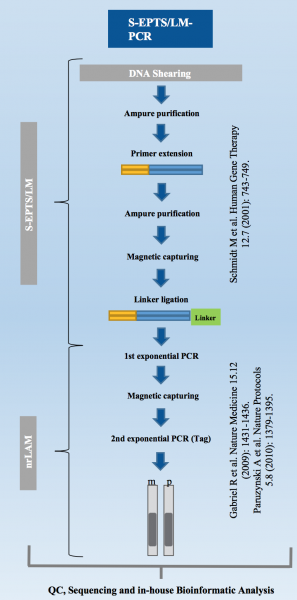
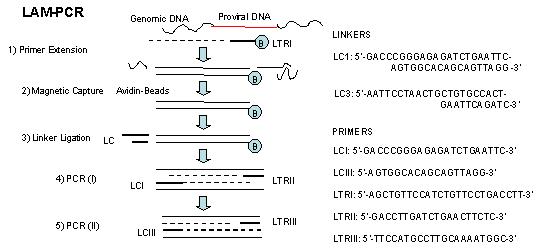
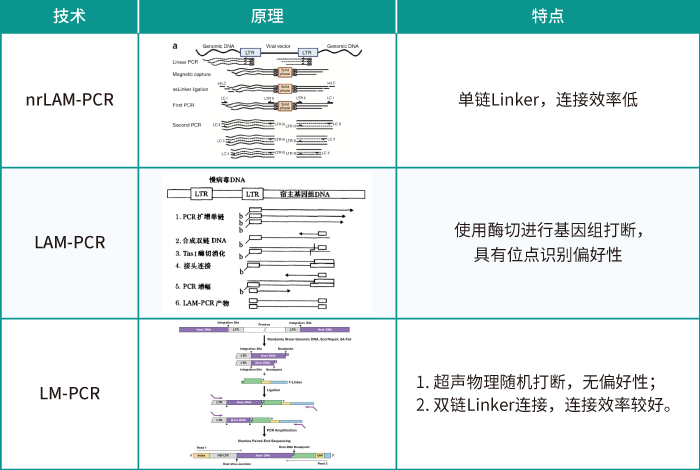
LAM-HGTGTS (Linear Amplification-mediated high-throughput genome-wide translocation sequencing) Our Working Protocol.

LAM-PCR workflow and the corresponding adLIMS automated process. The... | Download Scientific Diagram

Analysis of polyclonal vector integration sites using Nanopore sequencing as a scalable, cost-effective platform | bioRxiv

Schematic of AAV-AAV junctions isolated following LAM-PCR. A complete... | Download Scientific Diagram
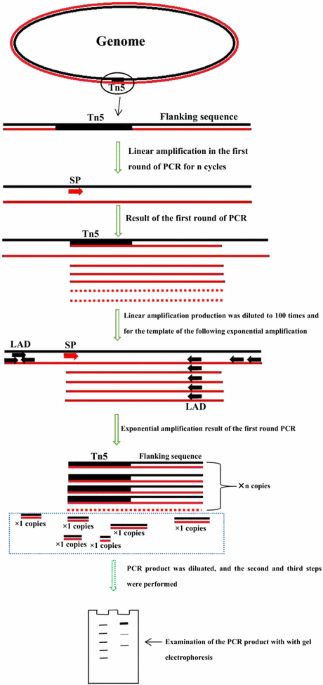
Linear and exponential TAIL-PCR: a method for efficient and quick amplification of flanking sequences adjacent to Tn5 transposon insertion sites | AMB Express | Full Text

Linear Amplification Mediated PCR – Localization of Genetic Elements and Characterization of Unknown Flanking DNA | Protocol (Translated to Italian)

Linear Amplification Mediated PCR – Localization of Genetic Elements and Characterization of Unknown Flanking DNA | Protocol (Translated to Italian)
HaSAPPy: A tool for candidate identification in pooled forward genetic screens of haploid mammalian cells | PLOS Computational Biology

VISPA: a computational pipeline for the identification and analysis of genomic vector integration sites | Genome Medicine | Full Text

LAM-PCR. a, Outline of the LAM-PCR method. 1, linear PCR by repeated... | Download Scientific Diagram

Schematic of a linear amplification-mediated polymerase chain reaction... | Download Scientific Diagram

Outline of linear amplification-mediated (LAM)-PCR. A new combination... | Download Scientific Diagram
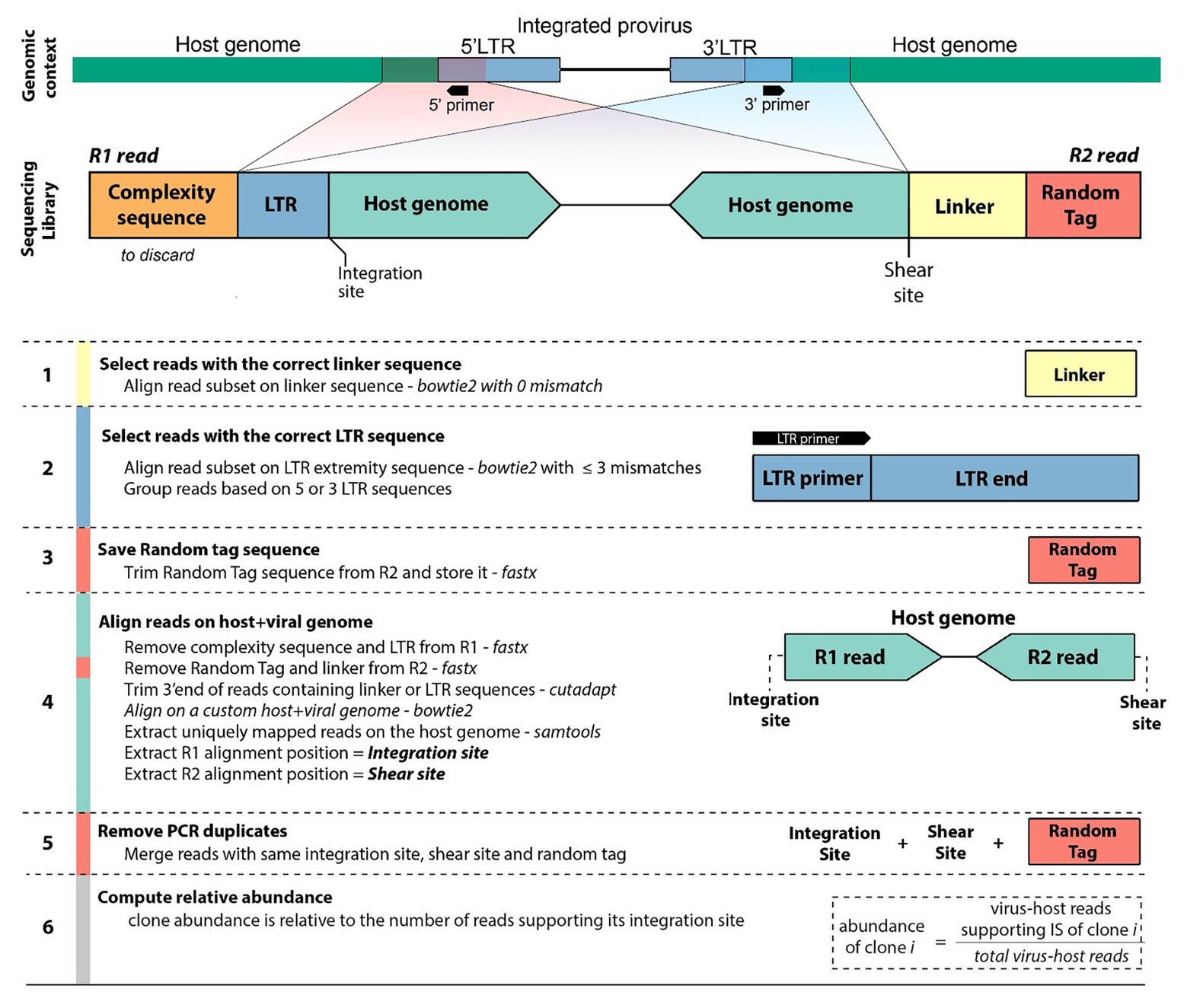
Frontiers | An Improved Sequencing-Based Bioinformatics Pipeline to Track the Distribution and Clonal Architecture of Proviral Integration Sites

Tagmentation-based analysis reveals the clonal behavior of CAR-T cells in association with lentivector integration sites - ScienceDirect

Integration Mapping of piggyBac-Mediated CD19 Chimeric Antigen Receptor T Cells Analyzed by Novel Tagmentation-Assisted PCR - eBioMedicine